Adhemarius (Clark, 1916)
|
publication ID |
https://doi.org/10.1093/isd/ixz025 |
|
persistent identifier |
https://treatment.plazi.org/id/038A581A-3A40-1C06-FF22-F8C8FC27D3A3 |
|
treatment provided by |
Felipe |
|
scientific name |
Adhemarius |
| status |
|
Classification of Adhemarius View in CoL , Orecta , and Trogolegnum
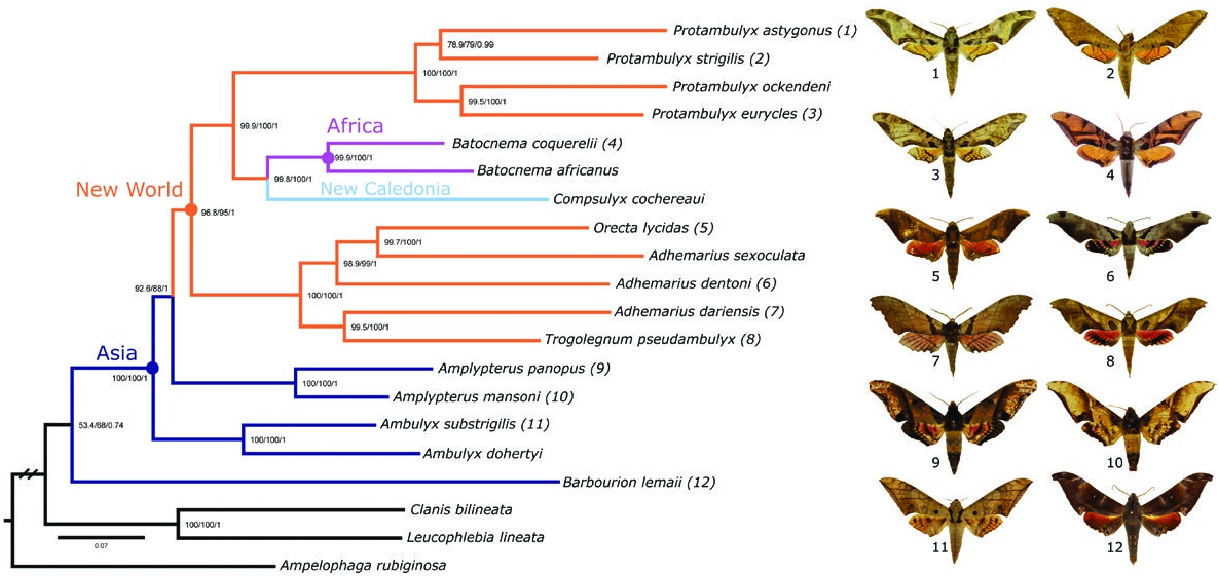
With regard to the phylogenetic relationships of Adhemarius , Orecta , and Trogolegnum , the present study ( Fig. 2 View Fig ) found a topology in which the Adhemarius donysa -group + Trogolegnum split off first, followed by the Adhemarius gannascus -group, leaving a terminal sister-group pairing of the A. sexoculata -group and Orecta . This result contrasts with Cardoso (2015), who found different patterns of relationship among these groups, depending upon the analytical method and data set used: in their IW MP analysis ( Cardoso 2015: Fig. 1.6 View Fig ), Orecta is first to branch off, followed by the sexoculata -group, then the gannascus -group, then finally Trogolegnum as sister to the donysa -group. In contrast, in the results of their BI analysis ( Cardoso 2015: Fig. 1.7 View Fig ), the sexoculata -group branched off first, followed by a trichotomy comprising Orecta , the gannascus -group and the donysa -group + Trogolegnum . However, all analyses agree that T. pseudambulyx is simply a member of the A. donysa species-group, albeit one with a reduced proboscis and labial palps, and, like Orecta , spinulose abdominal tergites and nonspinose abdominal sternites ( Rothschild and Jordan 1903). However, the phylogenetic relationships of Orecta , also considered by Rothschild and Jordan (1903) to be a derivative of Adhemarius , remain obscure, with each of the three analyses suggesting a different placement. We therefore consider it premature to make any formal changes to the classification of the three genera. In addition, although most of the relationships were recovered with high support, we should point out that mitochondrial genomes are maternally inherited, can introgress between hybridizing species and that the genes in the mitochondrial genome are tightly linked ( Avise and Ellis 1986). It is therefore possible that the trees obtained here merely represent a deviating gene history and not the actual evolutionary history of the species involved ( Ballard 2000). Phylogenomic studies currently in progress, which focus on the nuclear genome using anchored hybrid enrichment (Kawahara et al. in preparation) and ultra-conserved elements (Rougerie et al. in preparation), will show whether there is any discrepancy between the mitochondrial and nuclear genomes and are expected to unambiguously resolve the placement and relationships of ambulycine taxa, finally allowing taxonomic decisions to be made.
No known copyright restrictions apply. See Agosti, D., Egloff, W., 2009. Taxonomic information exchange and copyright: the Plazi approach. BMC Research Notes 2009, 2:53 for further explanation.